Assignment 5
Due date: Thursday, October 19th at midnight (Thursday night).
0) Be sure to use version control ("git"), as you develop your code. We suggest you create a new directory to hold this assignment, "assignment5" for example, and initialize a new git repository within it.
$ mkdir assignment5
$ cd assignment5
$ git init
git add ...., git commit" repeatedly as you add to your scripts. You will hand in the output of "git log" for your assignment repository as part of the assignment. You must have a significant number of commits representing the modifications, alterations and changes in your scripts. If your log does not show a significant number of meaningful commits you will loose marks.
This assignment will explore the Capture-recapture technique of estimating wildlife populations. Suppose we are estimating the population of a specific species of fish in a lake. This technique involves capturing a sample of the fish, tagging the fish so that they now have markers, releasing the fish back into the lake, and then, once the tagged fish have sufficiently spread thoughout the lake, returning to the lake and capturing a second sample of fish. Based on the number of previously-tagged fish in the second sample, the population of the fish in the lake can be estimated.
We will examine a variation on this technique known as inverse sampling capture-recapture. Let us assume that there are \(N\) fish in a given lake, and we have tagged \(M\) of them. This technique estimates \(N\) by counting the number of fish that need to be caught to catch \(m\) previously-tagged fish. If the number of fish that need to be caught is \(Y\), then \(N\) can be estimated using the equation $$\hat{N} = \frac{MY}{m}$$
We will perform a simulation of this type of population estimation, and examine the distribution of the estimates of the population of fish.
1) Create a file named Capture.Recapture.Utilities.R. It will contain the functions described in part 1 of the assignment. Note that, for the functions listed below, you will need to determine for yourself what arguments the functions need.
1a) Create a function called tag.population. This function will randomly pick \(M\) fish to be tagged out of the total population of \(N\) possible fish. The same fish may not be tagged twice. The function should return the indices of the tagged fish. The sample function might be useful here.
1b) Create a function called catch.fish. This function will randomly catch fish from the lake until \(m\) previously-tagged fish have been caught. Note that the fish are thrown back into the lake after each has been caught. Your function should use a population of tagged fish to determine if a tagged fish has been caught, not just a probability of catching a tagged fish. The function will return the number of fish that had to be caught to catch the \(m\) fish.
1c) Create a function called calc.N. This function will return an estimate for \(N\). Note that \(N\) should be an integer.
1d) Create a function called
simulate.pops. The function will perform the following steps:
- randomly generate a tagged fish population,
- calculate the number of fish it takes to catch \(m\) previously-tagged fish,
- estimate and return \(N\) using this data.
The function should be crafted so that it can be called using one of the *apply family of functions.
Your functions should behave similar to this:
>
> source("Capture.Recapture.Utilities.R")
>
> tagged.pop <- tag.population(100, 10)
>
> tagged.pop
[1] 54 59 84 49 24 33 64 74 50 41
>
> catch.fish(tagged.pop, 100, 5)
[1] 60
>
> calc.N(10, 60, 5)
[1] 120
>
2) Create an R script, named Run.Capture.Recapture.R, which:
- sources the file "Capture.Recapture.Utilities.R"
- reads three arguments from the command line. These arguments are \(N\), \(M\) and \(m\).
- The script should use one of the
*applyfamily of functions to runsimulate.pops1000 times with the values of \(N\), \(M\) and \(m\). - The script should create a normalized histogram of the estimated values of \(N\). Use the
histcommand to do this. Note that this will create a file calledRplots.pdfin your assignment directory. - If the command-line arguments are not numeric the script should exit with an appropriate error message.
- If the values of the command-line arguments do not make sense the script should exit with an appropriate error message.
- If the number of command-line arguments is not 3 the script should exit with an error message.
An example of your histogram, for \(N=1000\), \(M=100\) and \(m=100\), might look like this, where a distribution has been added to the plot (you do not need to add a distribution to your plot). This is an example of negative binomial distribution. 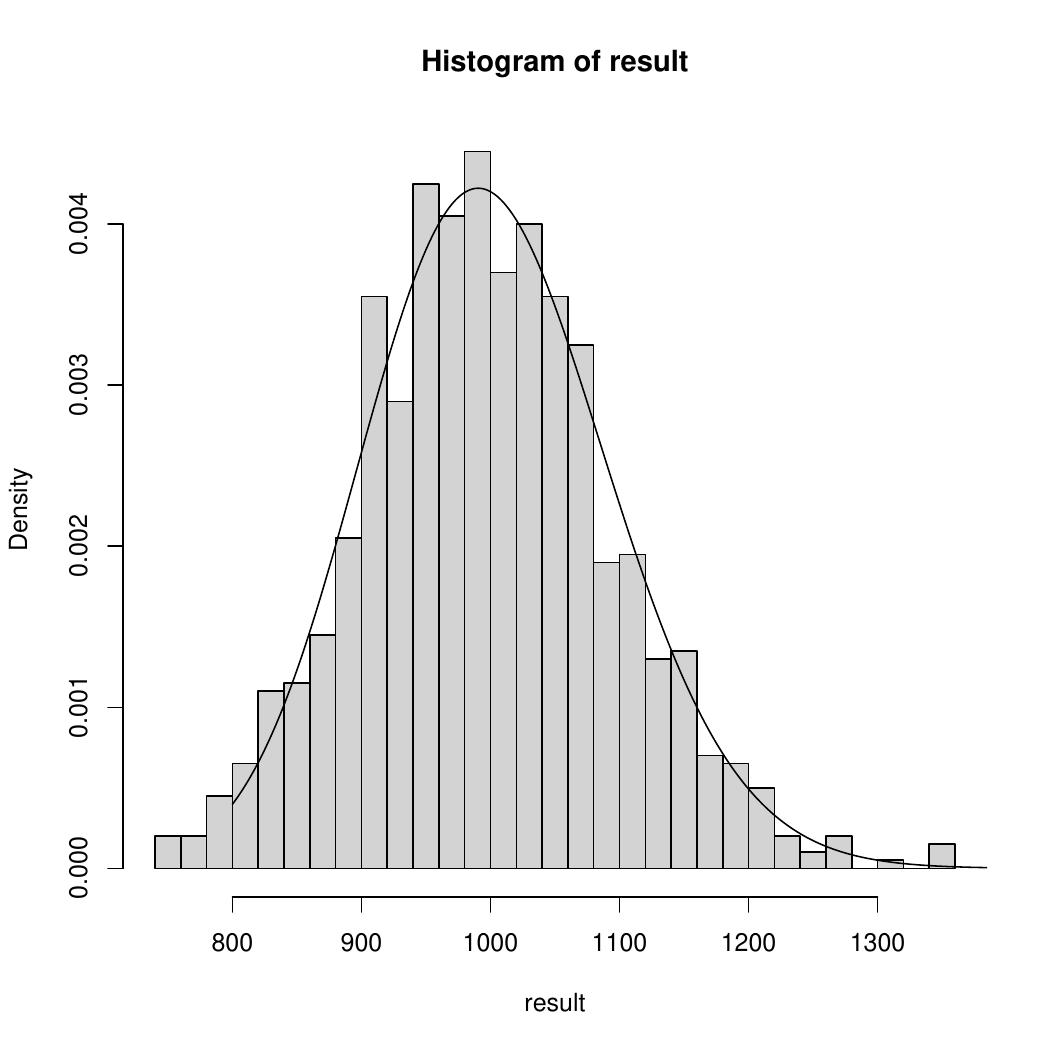
Note that, starting with this assignment and for the rest of the semester, you will be expected to use coding best practices in all of the work that you submit. This includes, but is not limited to:
- Plenty of comments in the code, describing what you have done.
- Sensible variable names.
- Explicitly returning values, if the function in question is returning a value.
- Not using the print() function to return values.
- Proper indentation of code blocks.
- No use of global variables.
- Using existing R functionality, when possible.
- Creating modular code. Using functions.
- Never copy-and-pasting code!
Submit your Capture.Recapture.Utilities.R, Run.Capture.Recapture.R and Rplot.pdf files and the output of "git log" from your assignment repository.
Both R scripts must be added and committed frequently to the repository. To capture the output of 'git log' use redirection ( git log > git.log, and hand in the "git.log" file).
Assignments will be graded on a 10 point basis. Due date is October 19th 2023 (midnight), with 0.5 point penalty per day for late submission until the cut-off date of October 26th 2023, at 9:00am.
